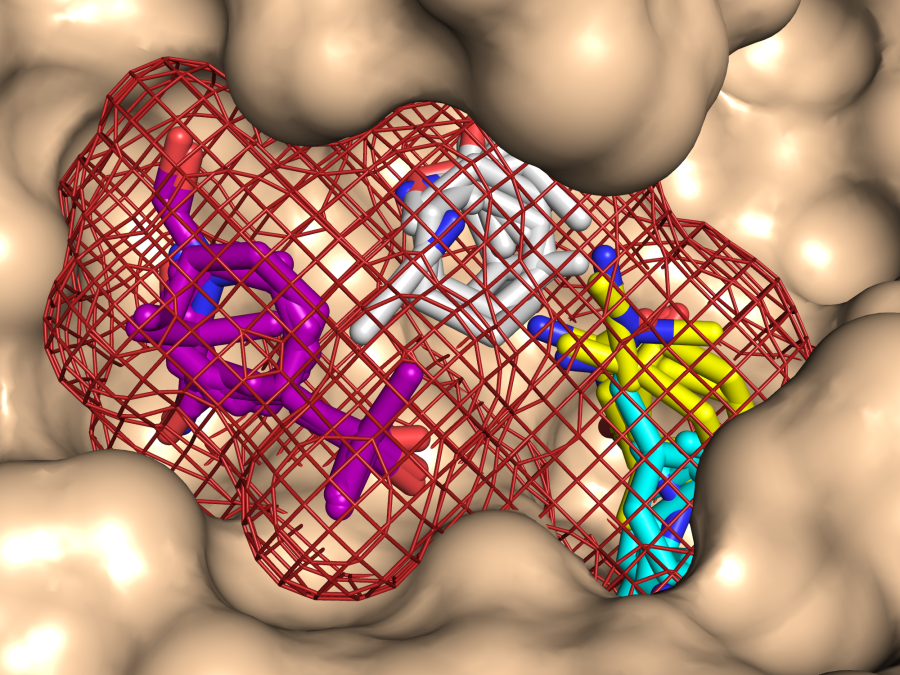
Target the right proteins with the right molecules. Computational solvent mapping identifies druggable binding sites and classifies what type of molecules they will bind.

Model protein-protein complexes with our protein-protein docking software, which is consistently among the best performers in CAPRI and has been used by thousands of researchers all over the world.

Engage with our scientists to customize our software and receive modeling services. Tailored software and services provides your company with the modeling you need to solve your problems.
